Research Agenda
Our research is focused on cells and molecules of the immune system. We would like to understand the mechanistic details of how these cells survive, signal and proliferate. We study receptors on the membranes of these cells as well as transcription factors in their nuclei. We are actively pursuing to develop our knowhow about cellular and molecular biology to marketable applications.
Our current research projects aim to a) model genetically inherited rare diseases by using CRISPR/Cas9 genome editing b) develop nanobody based bio-reagents targeting signaling pathways in lymphocytes and c) identify rules of dimerization of human transcription factors. Students working on these projects will be exposed to basic scientific research in immunology and molecular biology.
Our research is focused on cells and molecules of the immune system. We would like to understand the mechanistic details of how these cells survive, signal and proliferate. We study receptors on the membranes of these cells as well as transcription factors in their nuclei. We are actively pursuing to develop our knowhow about cellular and molecular biology to marketable applications.
Our current research projects aim to a) model genetically inherited rare diseases by using CRISPR/Cas9 genome editing b) develop nanobody based bio-reagents targeting signaling pathways in lymphocytes and c) identify rules of dimerization of human transcription factors. Students working on these projects will be exposed to basic scientific research in immunology and molecular biology.
The T Lymphocyte Receptor
T lymphocytes are essential for the functioning of the adaptive immune system. This is demonstrated by genetic or virus induced human immunodeficiencies resulting from a lack of T-cells. T-cells recognize changes in their environment through a specific T-cell receptor (TCR) protein expressed on their surface. This receptor binds to peptide antigens presented in the context of major histocompatibility complex (MHC) proteins on the surface of antigen presenting cells. TCR binding to its ligand causes signaling that re-programs the cell by turning on new genes. TCR first gets expressed on the surface of T cell precursors that also have two co-receptors on their surface named CD4 and CD8. These double positive (DP) thymocytes have the business of generating all the T lymphocytes in the body. Re-programming by new gene expression is important for the development of the thymocyte precursors of T cells in the thymus and for the response of T lymphocytes in peripheral lymphoid organs. We are interested in the structure of the TCR on the surface of T lymphocytes, the cytoplasmic signaling proteins that convey the signal to the nucleus and how new genes get turned on by TCR signals.
T lymphocytes are essential for the functioning of the adaptive immune system. This is demonstrated by genetic or virus induced human immunodeficiencies resulting from a lack of T-cells. T-cells recognize changes in their environment through a specific T-cell receptor (TCR) protein expressed on their surface. This receptor binds to peptide antigens presented in the context of major histocompatibility complex (MHC) proteins on the surface of antigen presenting cells. TCR binding to its ligand causes signaling that re-programs the cell by turning on new genes. TCR first gets expressed on the surface of T cell precursors that also have two co-receptors on their surface named CD4 and CD8. These double positive (DP) thymocytes have the business of generating all the T lymphocytes in the body. Re-programming by new gene expression is important for the development of the thymocyte precursors of T cells in the thymus and for the response of T lymphocytes in peripheral lymphoid organs. We are interested in the structure of the TCR on the surface of T lymphocytes, the cytoplasmic signaling proteins that convey the signal to the nucleus and how new genes get turned on by TCR signals.
T Lymphocytes and Natural Killer cells
Signaling by the TCR activates various transcription factors that translocate to the nucleus to bind to transcriptional control elements of TCR responsive genes. NF-AT, and NF-kappaB, are two such transcription factors that wait in an inactive form in the cytoplasm before signaling happens. Nuclear translocation of these factors results from cytoplasmic phosphorylation and dephosphorylation events. If non-mamalian species have signaling proteins that are similar to but not identical to NF-AT, and NF-kappaB activating proteins, we may be able to introduce these factors into mammalian cells and modulate TCR signal transduction. We generated reporter cell lines that express green fluorescent protein (GFP) in response to NF-AT, and NF-kappaB activation. In past experiments we performed yeast two hybrid screens to identify novel interactors of the CD3 delta subunit of the TCR. In collaboration with Tolga Sutlu and Adil Doganay Duru, we transferred all components of the TCR complex into Natural Killer cells, reprogramming them to be antigen specific killers of melanoma. This technology is being commercially developed as an immunotherapy for cancer patients. How cytotoxic cells kill targets, mechanisms of activation, degranulation and serial triggering are current areas of interest.
Signaling by the TCR activates various transcription factors that translocate to the nucleus to bind to transcriptional control elements of TCR responsive genes. NF-AT, and NF-kappaB, are two such transcription factors that wait in an inactive form in the cytoplasm before signaling happens. Nuclear translocation of these factors results from cytoplasmic phosphorylation and dephosphorylation events. If non-mamalian species have signaling proteins that are similar to but not identical to NF-AT, and NF-kappaB activating proteins, we may be able to introduce these factors into mammalian cells and modulate TCR signal transduction. We generated reporter cell lines that express green fluorescent protein (GFP) in response to NF-AT, and NF-kappaB activation. In past experiments we performed yeast two hybrid screens to identify novel interactors of the CD3 delta subunit of the TCR. In collaboration with Tolga Sutlu and Adil Doganay Duru, we transferred all components of the TCR complex into Natural Killer cells, reprogramming them to be antigen specific killers of melanoma. This technology is being commercially developed as an immunotherapy for cancer patients. How cytotoxic cells kill targets, mechanisms of activation, degranulation and serial triggering are current areas of interest.
|
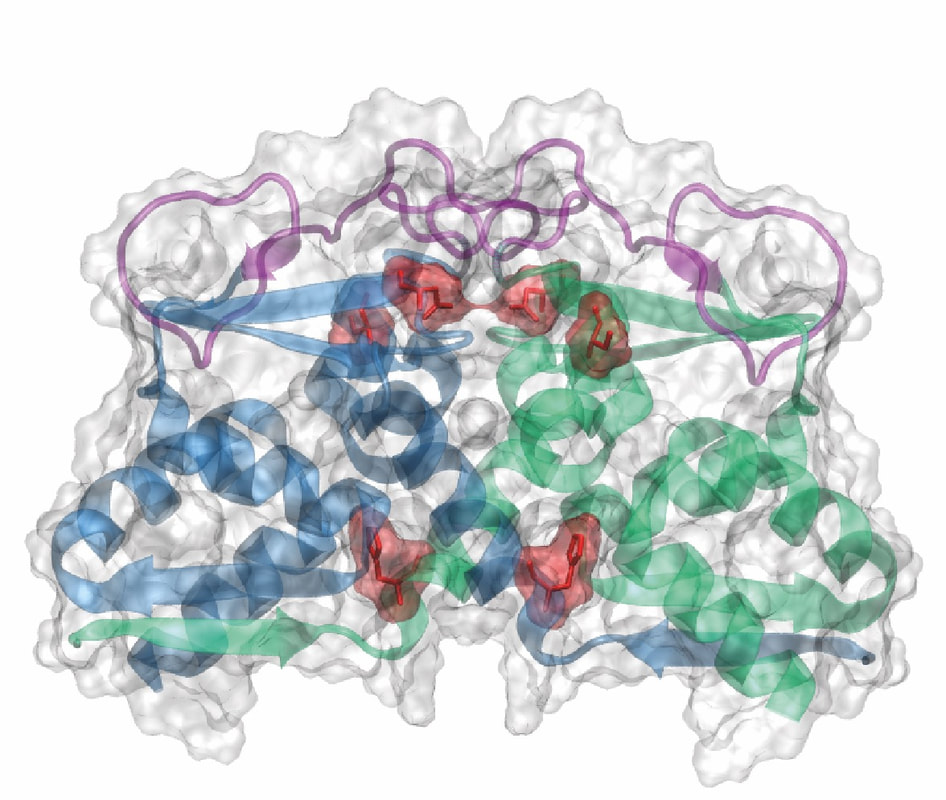
A model of the BTB domain of the PATZ-1 transcription factor which is based on our X-ray crystal structure of the protein.
|
The Interleukin 7 Receptor
Signaling by the receptor for the pro-survival cytokine, IL-7 (IL-7R) is critical for lymphocyte survival as demonstrated by immunodeficiencies observed in IL-7R gene knockout mice. IL-7 receptor is expressed in T lymphocytes as well as B lymphocytes. In humans, X-linked severe combined immunodeficiency (X-SCID) syndrome develops when a gene encoding a component of the IL7R signaling complex (the cytokine receptor common gama chain) is mutated on the X chromosome. The transcription regulatory regions of the IL7-R gene are not clearly defined, and whether the same transcription factors activate the transcription of this gene in these different lineage cells is not known. We identified transcriptional enhancers of the IL-7R gene locus using bioinformatics and gene reporter assays. We modified the IL-7R gene locus by “recombineering” bacterial artificial chromosomes (BAC) that we re-introduce into tissue culture cell lines that express IL7-R and into transgenic mice. |
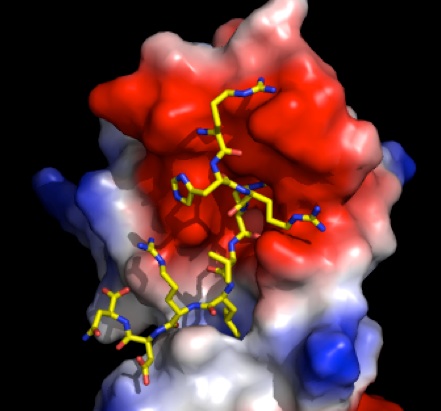
PATZ-1 protein, our favorite transcription factor
Recently we dove into the field of epigenetics and cancer biology! How transcription factors change the histone code is an exciting field. We work with zinc-finger transcription factors that we think bind to methylated histones and the tumor suppressor protein p53. We are purifying these transcription factor proteins by expressing them in E. coli for structural and functional studies. Patz-1 is a zinc finger protein that binds to p53 and inhibits its function. We use cutting edge techniques such as whole genome RNA Sequencing to identify the pathways regulated by Patz-1 and its role during the p53 dependent DNA damage response. Sofia Piepoli recently crystallised and solved the structure of the BTB domain of this protein. BTB domains are protein-protein interaction domains through which these transcription factors form homo- or heterodimers. These dimers form landing pads for epigenetic co-repressor proteins. We are trying to solve the code with which dimerization is regulated.
Recently we dove into the field of epigenetics and cancer biology! How transcription factors change the histone code is an exciting field. We work with zinc-finger transcription factors that we think bind to methylated histones and the tumor suppressor protein p53. We are purifying these transcription factor proteins by expressing them in E. coli for structural and functional studies. Patz-1 is a zinc finger protein that binds to p53 and inhibits its function. We use cutting edge techniques such as whole genome RNA Sequencing to identify the pathways regulated by Patz-1 and its role during the p53 dependent DNA damage response. Sofia Piepoli recently crystallised and solved the structure of the BTB domain of this protein. BTB domains are protein-protein interaction domains through which these transcription factors form homo- or heterodimers. These dimers form landing pads for epigenetic co-repressor proteins. We are trying to solve the code with which dimerization is regulated.
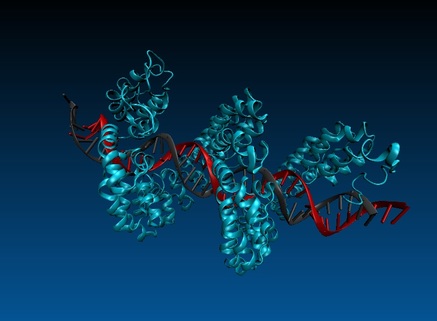
Genome editing technologies
Recently we established novel genome editing technologies in our lab. We use Transcription activator like effector nucleases (TALENs) and the CRISPR/Cas9 system to generate genomic mutations in tissue culture cell lines. We have generated numerous tissue culture cell line derivatives to perform structure-function studies. One of our targets is the IL-7R gene locus. We study transcription factor binding sites in the transcriptional enhancers of re-engineered mutant IL-7R loci to identify the function of these gene regulatory regions. Another target is the p53 gene. Our mutant cell lines allowed us to discover new mechanisms of control of this tumor suppressor, involving alternative splicing regulated by putative exonic splicing enhancer sequences.
Recently we established novel genome editing technologies in our lab. We use Transcription activator like effector nucleases (TALENs) and the CRISPR/Cas9 system to generate genomic mutations in tissue culture cell lines. We have generated numerous tissue culture cell line derivatives to perform structure-function studies. One of our targets is the IL-7R gene locus. We study transcription factor binding sites in the transcriptional enhancers of re-engineered mutant IL-7R loci to identify the function of these gene regulatory regions. Another target is the p53 gene. Our mutant cell lines allowed us to discover new mechanisms of control of this tumor suppressor, involving alternative splicing regulated by putative exonic splicing enhancer sequences.
Induced pluripotent stem cell (iPSC) technologies
To capitalize on our expertise in TALEN and CRISPR/Cas9 genome editing technologies, we have initiated several projects to modify the genome of iPSC generated from fibroblasts of human patients. We aim to perform gene therapy on these stem cells by fixing known mutations using our genome editing tools. Two main targets are genes that cause a) genetic cardiac arrhythmias and b) immuno-deficiencies. A high percentage of consanguineous intermarriage in Turkey results in a relatively high frequency of recessive genetic diseases in the population. We are obtaining skin biopsies from Turkish patients and generating iPSC, on which we perform genome editing using our CRISPR/Cas9 tools.
To capitalize on our expertise in TALEN and CRISPR/Cas9 genome editing technologies, we have initiated several projects to modify the genome of iPSC generated from fibroblasts of human patients. We aim to perform gene therapy on these stem cells by fixing known mutations using our genome editing tools. Two main targets are genes that cause a) genetic cardiac arrhythmias and b) immuno-deficiencies. A high percentage of consanguineous intermarriage in Turkey results in a relatively high frequency of recessive genetic diseases in the population. We are obtaining skin biopsies from Turkish patients and generating iPSC, on which we perform genome editing using our CRISPR/Cas9 tools.
|
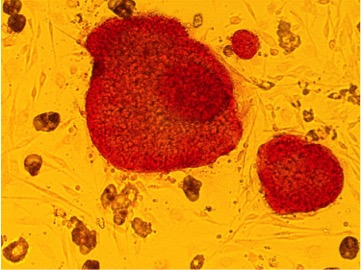
An iPSC colony generated in the laboratory.
|
Anti-cancer drugs
Our work on p53 has led us to design and screen small molecule inhibitors which can be used as anti-proliferative agents against cancer. We have targeted the molecular interaction surface between the MDM2 and p53 proteins. We have several candidate molecules that disrupt this interaction and lead to the accumulation of the p53 protein and result in the death of cancer cells. This activity is specific as p53 deficient cancer cells are not affected by these compounds. We are presently pursuing IP disclosure discussions to develop these compounds as marketable anti-proliferative drugs. More recently we generated nanobodies that target the interaction between p53 and MDM4 and are exploring computational and experimental methods to make these nanobodies cell-permeable. This will allow us to express and purify these nanobodies targeting intracellular interactions and develop them as in vivo applicable drugs. |
Development of Drugs
We are so excited to participate in an industrial project in collaboration with Dr. Tolga Sutlu's research group and ILKO Pharmaceuticals that aims to develop biosimilar drugs. What are biosimilars? These are generally antibody drugs that are proven to be active in patients whose patent protection is or will be expiring. Our aim is to develop the necessary laboratory infrastructure to a) grow chinese hamster ovary (CHO) tissue culture cell lines that produce an antibody in bioreactors and b) analyze the structure and function of this antibody using equipment such as a flow cytometer, Biacore surface plasmon resonance (SPR) and mass spectrophotometry. This project has concluded but we also have close collaborations with Turkish pharmaceutical companies such as Kocak Pharma and Turgut Ilaclari to develop novel drugs. An active project in the laboratory is the develepment of a biosimilar version of Anakinra, a protein-drug that is a version of the Interleukin-1 Receptor Antagonist that is effective in anti-inflammatory signaling. We are developing nanobodies that can target the same receptor as well as IL1Ra expressed in the periplasmic space of E.coli. This effort led us to discover novel rules governing protein expression in prokaryotic systems. We are developing optimized signal peptides and modifying cellular pathways that allow maximal protein expression in the bacterial periplasm.
We are so excited to participate in an industrial project in collaboration with Dr. Tolga Sutlu's research group and ILKO Pharmaceuticals that aims to develop biosimilar drugs. What are biosimilars? These are generally antibody drugs that are proven to be active in patients whose patent protection is or will be expiring. Our aim is to develop the necessary laboratory infrastructure to a) grow chinese hamster ovary (CHO) tissue culture cell lines that produce an antibody in bioreactors and b) analyze the structure and function of this antibody using equipment such as a flow cytometer, Biacore surface plasmon resonance (SPR) and mass spectrophotometry. This project has concluded but we also have close collaborations with Turkish pharmaceutical companies such as Kocak Pharma and Turgut Ilaclari to develop novel drugs. An active project in the laboratory is the develepment of a biosimilar version of Anakinra, a protein-drug that is a version of the Interleukin-1 Receptor Antagonist that is effective in anti-inflammatory signaling. We are developing nanobodies that can target the same receptor as well as IL1Ra expressed in the periplasmic space of E.coli. This effort led us to discover novel rules governing protein expression in prokaryotic systems. We are developing optimized signal peptides and modifying cellular pathways that allow maximal protein expression in the bacterial periplasm.
The joint Sutlu-Erman research teams participated in close harmony on the biosimilar project, along with our colleagues from ILKO Pharmaceuticals and our acting president. Didem, Nazife, Melike, Tolga, Hakan, Hatice Hn, Ronay, Ayse Hn., Mert, Duygu Hn., Alper Tunga, Batu, Lola, Ayhan, Pegah, Cevriye, Sinem, Sofia.
LincRNAs
We are very excited by long intergenic non-coding RNAs. Recent developments in this field demonstrate that many biological phenomena including transcription activation, epigenetics, development, signaling and the stress response involve these molecules. We are actively re-thinking all of the processes we study with a lincRNA viewpoint! The genome reveals endless information. There is no "junk DNA", estimates of gene numbers have to be re-evaluated. Is there such a thing as a pseudogene? Is there a hidden RNA world? These are exciting times for biology.
We are very excited by long intergenic non-coding RNAs. Recent developments in this field demonstrate that many biological phenomena including transcription activation, epigenetics, development, signaling and the stress response involve these molecules. We are actively re-thinking all of the processes we study with a lincRNA viewpoint! The genome reveals endless information. There is no "junk DNA", estimates of gene numbers have to be re-evaluated. Is there such a thing as a pseudogene? Is there a hidden RNA world? These are exciting times for biology.